Identification Analysis by Neural Networks
Introduction
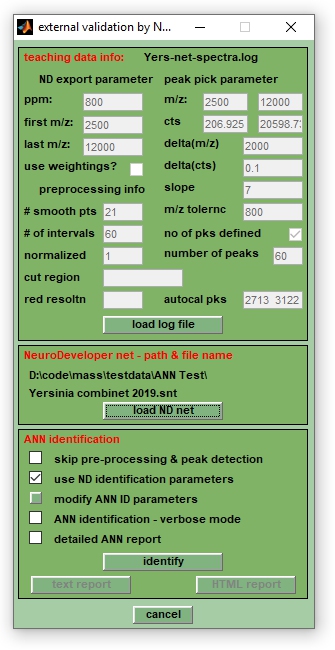
The function Identification by Neural Networks is basically an interface between MicrobeMS and Synthon's NeuroDeveloper™, a software for teaching and validating ANN models with microbial mass spectra. Based on pre-defined neural network models, the interface can be used to classify mass spectra into pre-defined categories (i.e. microbial genera, species, subspecies, etc.). Spectral pre-processing, features selection and ANN classification of a priori unknown spectra is automatically performed. This is achieved by utilizing the runtime environment of the NeuroDeveloper™ software which does not require a software license from Synthon. Spectral data can be therefore classified without the NeuroDeveloper™ software on the basis of predefined network libraries. The NeuroDeveloper™ is, however, required if you wish to create and validate own ANN models.
Synthon GmbH contact address: Analytics and Pattern Recognition Im Neuenheimer Feld 69120 Heidelberg GERMANY phone: +49 6221 50 257 900 fax: +49 6221 50 257 909 email: info@synthon-analytics.com internet: http://www.synthon-analytics.de
Some classes of artificial neural networks (ANN) are computational models which are popular means to model complex relationships between inputs and outputs and to find patterns. In supervised learning so called multilayer-perceptron (MLP) ANNs require a set of training samples which can be used to infer a classifier for predicting class assignments. To avoid overfitting and to assess the robustness of ANN class assignment, the general strategy of analysis by MLP-ANNs routinely includes procedures of training, internal validation and (external) testing, ideally under blinded conditions. Training and internal validation requires samples with of known class assignments (i.e. labeled spectra); the model performance can be then determined on the basis of errors between the obtained outputs and desired target values of both, the training and the internal validation subsets. At the training stage ANN performance can be optimized by modifying the way of pre-processing, adding or eliminating spectral features, or changing the network's architecture. When training is finished the classifier can be challenged by an external validation (test) subset.
Create Neurodeveloper™ ANN models
Of note, a license of the NeuroDeveloper™ software will be required.
Compilation of mass spectra subsets for teaching and internal validation
1. Load the respective MALDI-TOF mass spectra
2. Perform spectral pre-processing. All spectra used for teaching and internal validation are required
to be processed in exactly the same way by utilizing the same sequence of pre-processing steps and
identical pre-processing parameters
3. Run the peak detection routine. Again the same parameters should be applied to all mass spectra,
see function peak detection for details
4. Assign mass spectra to individual classes by using the function class assignment
5. Export the mass spectra by the function Store peaklist data in a format specific to the NeuroDeveloper™ software (export to NeuroDeveloper™). When exporting spectra a log file will be automatically created that is required for ANN classification analysis (see below)
6. Train and validate the ANN model by the NeuroDeveloper™ software by using the exported spectra files.
Store the network model (*.snt data format). For details please refer to the NeuroDeveloper™ software
manual
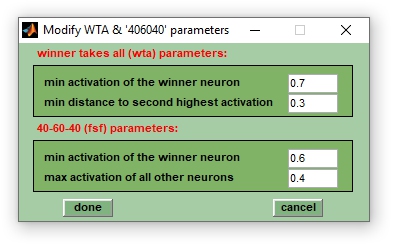
Classification
to be continued (Dec 2022)
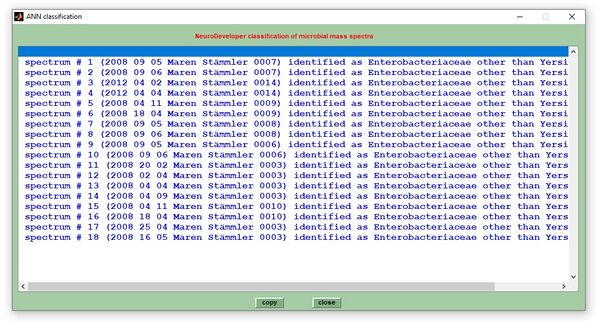
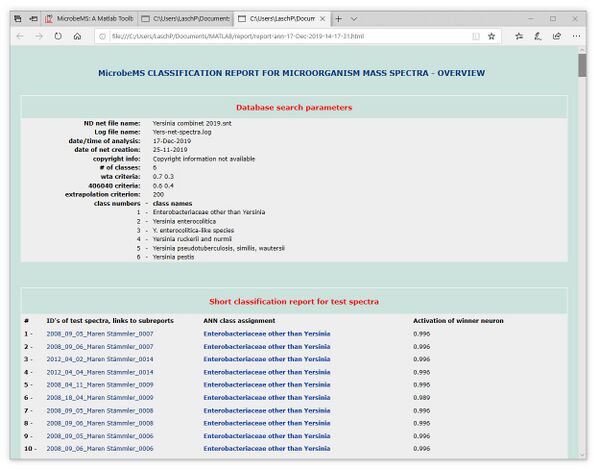
Classification reports
to be continued (Dec 2022)