Export XML Data
MicrobeMS facilitates the generation of XML data files from MALDI-ToF mass spectra, which are necessary for the utilization of MicrobeNet.
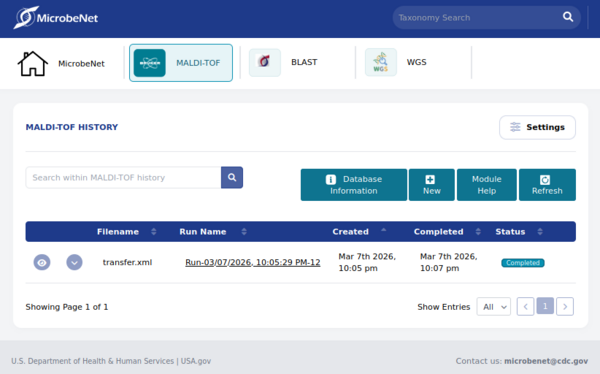
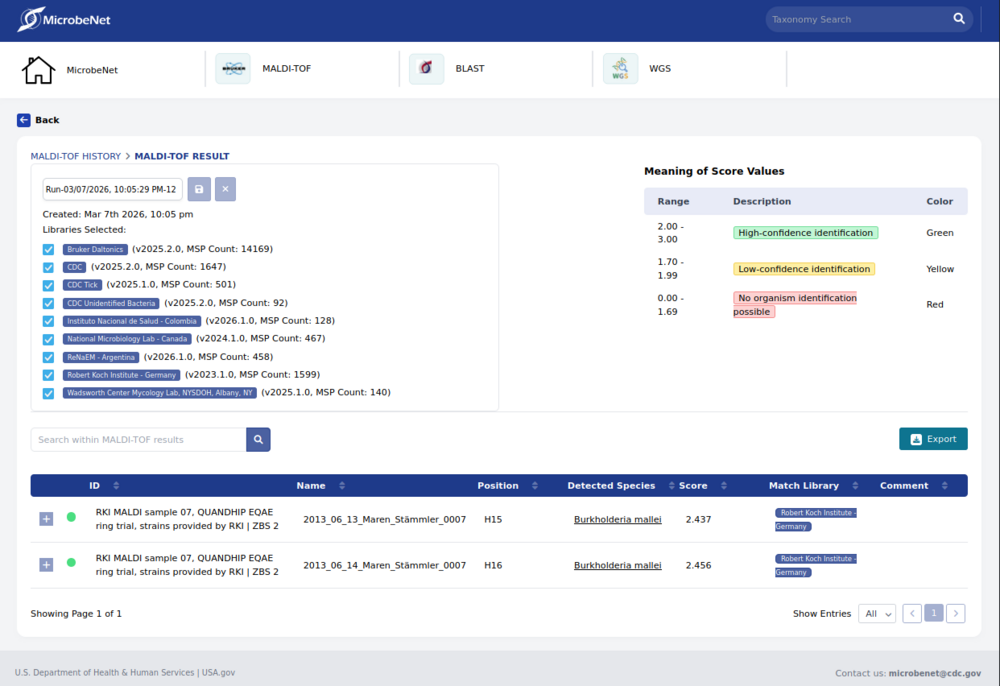
MicrobeNet is an free online resource maintained by the Centers for Disease Control and Prevention (CDC, Atlanta, USA). It is designed to assist microbiologists with identifying bacterial and fungal pathogens by MALDI-ToF MS and 16S/ITS gene sequencing (A WGS analysis module is planned). Users of the MicrobeNet MALDI module have the capability of submitting MALDI-ToF data for querying against Bruker Daltonics' MALDI Biotyper (MBT) pathogen database, databases maintained by the CDC (bacteria, unidentified bacteria, tick database) and databases of collaborating institutions like ReNaEM (Buenos Aires, Argentina), National Microbiology Lab (Winnipeg, Canada), Robert Koch-Institute (Berlin, Germany) and the Instituto Nacional de Salud (Bogota, Colombia). The objective of MicrobeNet is to identify rare and unusual pathogens online, thereby saving clinical laboratories time and money by eliminating the need to send specimens to specialized laboratories for identification.
XML files are typically generated automatically during spectrum measurement using Bruker MBT software. To accomplish this, the MBT software must be equipped with a patch. Further details on this process can be found on the CDC website. While it is theoretically possible to generate an XML file retrospectively using MBT's onboard tools from existing MALDI-ToF mass spectra, doing so is very labor-intensive (as of December 2025).
An XML MALDI file does not contain complete spectrum data but rather a peak table aside from some basic metadata. The Bruker MBT software generates peak data using predefined parameters. This peak table data is then stored in the XML file.
MicrobeMS' export XML data function enables direct conversion of MALDI spectrum data into the XML format. By this function MALDI-ToF mass spectrum data is automatically converted into a peak table format using predefined parameters for preprocessing and peak detection. The corresponding parameters can be viewed and adjusted in the file microbems.opt. Note that the MBT software and MicrobeMS use different methods of preprocessing and peak detection. The peak tables they generate may therefore differ slightly. However, systematic tests have demonstrated that the specific score values of the identification analyses vary only marginally.
To convert MALDI-ToF spectrum data to XML format, the data must first be loaded. Once the spectra have been selected (i.e., marked), the XML export (CDC MicrobeNet) option can be selected from the File pulldown menu. A dialog box will open where you can enter the name and path of the file to be saved. After confirming, the spectrum data will be processed, and the peak tables will be stored in an XML file. This file can then be used for online identification analysis in MicrobeNet.