Two-samples t-Tests
Jump to navigation
Jump to search
Introduction
This function performs consecutively two-sample t-tests in segments of the MALDI-ToF mass spectra using peak table data as inputs. t-tests are useful for biomarker screening in ensembles of labeled MALDI-ToF mass spectra that exhibit certain degree of similarity. A two-sample t-test in a given m/z segment returns a test decision for the null hypothesis that the peak intensity data in classes I and II arise from independent random samples from normal distributions with equal means and variances. The alternative hypothesis is that the peak intensity data come from populations with unequal means.
See also t-test (Wikipedia)
Parameter of two-samples t-tests
- m/z range: lower and upper bounds of the m/z region in which the series of t-tests are to be performed
- α: significance level of the t-tests
- dx (ppm): a parameter defining the relative width and thus the number of the spectrum segments. A spectrum segment centered at position covers a m/z interval of an absolute width equaling . The lower and upper bounds of the spectrum segments are defined by (lower bound) and (upper bound), respectively. Consequently, a spectrum segment of width dx=1000 and centered at =2000 Th would be 2 Th wide with boundaries located at 1999 and 2001 Th
- intensity: defines if barcode spectra (checkbox unchecked) or peak weighting factors (checked) are utilized as test inputs
- show histogram: shows a histogram with test outputs (p-values, AUC, etc.) and provides also the mean, median and the standard deviation of the test variables
Performing t-test series
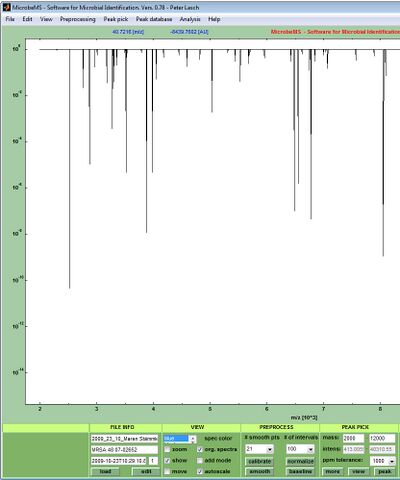
1. Load the mass spectral data files via the load spectra (Bruker data file format), import spectra from mzXML data, or the load MS multifile options of the File pull down menu.
2. Two-samples t-tests are carried out from labeled spectra, i.e. from spectra with a class assignment. To perform the test label two groups of spectra as class 1 and as class 2, respectively. Labeling, or class assignment, can be carried out by selecting the appropriate spectra and choosing class assignments → class X from the Edit pull down menu.
3. The t-test routine always starts from original MALDI-ToF mass spectra, i.e. spectral pre-processing and peak detection is carried out automatically using pre-defined parameters. Existing pre-processed spectra and pre-defined peak tables are ignored by the test routine.
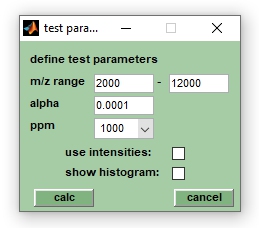
4. Define test parameter, such as α (significance level), the m/z range and dx (ppm) which has a default value of 1000 (relative, in ppm). The parameter dx defines the width of m/z segments in which spectra are divided during the test. Peaks found in the same m/z segment are considered identical while mass peaks in different segments are considered different peaks.
5. When finished select t-test from the Analysis pull down menu. Choose options plot decision for H(0), plot p-values or plot t-values, to obtain the respective outputs of the t-tests.
Output of the univariate t-test series
Example of the output from a serial t-test taken from the log file of MicrobeMS:
 that the peak intensity data in classes I and II arise from independent random samples from normal distributions with equal means and variances. The alternative hypothesis
that the peak intensity data in classes I and II arise from independent random samples from normal distributions with equal means and variances. The alternative hypothesis  is that the peak intensity data come from populations with unequal means.
is that the peak intensity data come from populations with unequal means.
 covers a m/z interval of an absolute width equaling
covers a m/z interval of an absolute width equaling  . The lower and upper bounds of the spectrum segments are defined by
. The lower and upper bounds of the spectrum segments are defined by ![{\displaystyle [{x_{i}*(1-dx)*0.5*10^{-6}}]}](https://wikimedia.org/api/rest_v1/media/math/render/png/e2ed469270c3f2578b386fb4df4eae02c9d3491d) (lower bound) and
(lower bound) and ![{\displaystyle [{x_{i}*(1+dx)*0.5*10^{-6}}]}](https://wikimedia.org/api/rest_v1/media/math/render/png/e1c7c01452ebbb7d73c9d7b9a035af06d7646d3b) (upper bound), respectively. Consequently, a spectrum segment of width dx=1000 and centered at
(upper bound), respectively. Consequently, a spectrum segment of width dx=1000 and centered at